DAMAGE-FREE SORTING
In recent years, researchers have noticed changes in their cells such as morphological changes, delayed cell growth, decrease in cell viability and alterations in gene expression after sorting. These post-sorting cell changes are a result of “sorter-induced cellular stress (SICS)” triggered by the traditional sorting techniques called Jet-in-air method, and therefore, sorting involving cells that are prone to SICS have been extremely challenging.
On-chip Sort has been used for wide variety of cell types, especially those that are too fragile to be sorted using other cell sorters, including neurons, iPS-derived cells, transfected cells, cardiomyocytes, hepatocytes, sperm cells, and adipocytes. Herein, we describe some of the results obtained in conjunction with our collaborators.
Effect of sorting on cell growth
In order to compare the effects of sorting on relatively robust HeLa cells between a conventional cell sorter (Jet-in-air method) and On-chip Sort (Flow shift method), 100 HeLa cells were sorted on both sorters. After sorting, collected cells were cultured in wells of a 96-well plate and the cell numbers were counted daily.
 Fig. 1. Growth curves of HeLa cells after sorting. The images were taken on day 3.
Fig. 1. Growth curves of HeLa cells after sorting. The images were taken on day 3.
Cells sorted by On-chip Sort began to grow two days after sorting, and cell number doubled on day 3 (Fig. 1). On the other hand, the cells sorted by conventional sorter only started growing three days after sorting, with a slower growth rate as compared to those sorted by On-chip Sort (Fig. 1).
Effect of sorting on cell morphology
The morphology of eosinophils that are highly susceptible to physical damage, were compared between a conventional sorter and On-chip Sort after sorting.
 Fig. 2. Morphology of peripheral blood eosinophils stained with hematoxylin and eosin, and fixed by Cytospin after sorting by On-chip Sort (left) and a conventional sorter (right).
Fig. 2. Morphology of peripheral blood eosinophils stained with hematoxylin and eosin, and fixed by Cytospin after sorting by On-chip Sort (left) and a conventional sorter (right).
Fig. 2 shows the morphologies of eosinophils from peripheral blood stained with hematoxylin and eosin, and fixed by CytoSpin after sorting by On-chip Sort (left) and a conventional sorter (right). Post-sorting morphological differences were clearly observed. Eosinophils sorted by On-chip Sort were intact, with cellular structures maintained. However, those sorted by conventional sorter were ruptured. It is speculated that sorting by conventional sorter has induced damages to the cell membranes.
Effect of sorting on RNA expressions
Many researchers sort cells into sub populations before analyzing the gene expression. In order to examine the effect of cell sorting on the difference in gene expression patterns between On-chip Sort and a conventional sorter, 50,000 Caco-2 and HeLa cells were sorted on both sorters. The change in gene expression level for each cell type was determined by microarray analysis using unsorted cells as control.

Fig. 3. Comparison of large changes in gene expression levels between sorted and unsorted HeLa cells (control) on (a) On-chip Sort and (b) conventional sorter A. In both (a) and (b), red dots represent up-regulated genes and green dots represent down-regulated genes. The numbers in percentages indicate percentages of all genes up-regulated (red) and down-regulated (green). (c) A histogram of unaffected gene expressions for Caco-2 and HeLa cells after sorting by On-chip Sort and conventional sorter A. The numbers in percentage indicate the fraction of unaffected genes over total genes for all categorized properties (apoptosis, cell adhesion, cell cycle, etc.) combined. The ratio of expression change is defined as the signal intensity of sorted cells/signal intensity of unsorted cells. The results are obtained in conjunction with Dr. Yasuda and Dr. Hagiwara from Cell Innovator Co., Ltd., and Dr. Tashiro from Kyushu University, Japan.
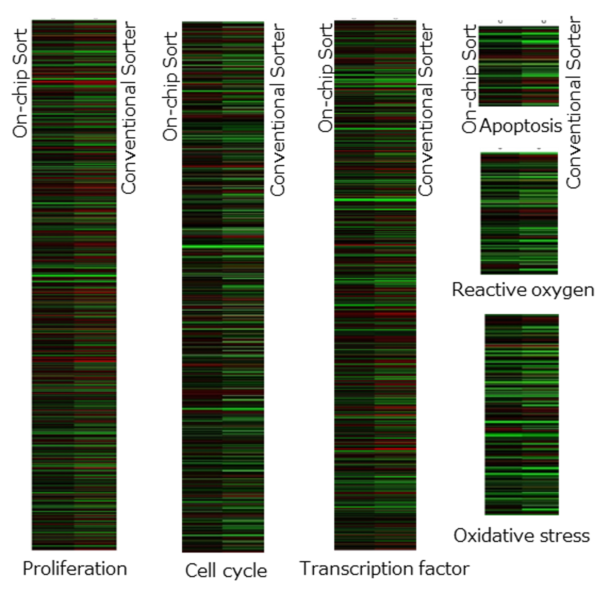
Fig. 4. Gene expression patterns of categorized properties with confirmed differences. In each properties, left and right columns represent data of On-chip Sort and conventional sorter, respectively. Black color indicates no change in gene expression, red color indicates up-regulation, and green down-regulation. The results are obtained in conjunction with Dr. Yasuda and Dr. Hagiwara from Cell Innovator Co., Ltd., and Dr. Tashiro from Kyushu University, Japan.
Fig. 3A and B show that HeLa cells sorted by On-chip Sort had less alteration in gene expression than those sorted by the conventional sorter. Furthermore, the number of genes with no change in gene expression level was 3-5 folds higher for On-Chip Sort than the conventional cell sorter (Fig. 3C). Fig. 4 shows the gene expression patterns of different properties. The properties displayed, namely proliferation, cell cycle, transcription factor, apoptosis, reactive oxygen and oxidative stress, were the ones that showed clear differences after sorting by On-chip Sort and conventional sorter. These differences in gene expression patterns between On-chip Sort and conventional sorter (Fig. 4) support the difference in growth rates between the two sorters shown in Fig. 1. Note that precautious measure must be taken when carrying out downstream analysis of sorted cells as transcription factors are likely to be affected. Although the gene expression analysis was done within 30 to 60 min after sorting for both types of cells, the result may differ depending on the post-sorting processes. Overall, differences in gene expression after sorting between On-chip Sort and conventional sorter can be observed.
Sorting of fragile cells
Neurons
In recent years, research on neural activity related to the pain and development for the visualization of pain have been carried out in various fields. The cell body of a sensory neuron (afferent neuron) called DRG neuron found in Dorsal Root Ganglions (DRGs) is responsible for the pain transmission from outside world and organs. However, there are many other types of cells that exist inside DRGs, and thus they consist of heterogeneous cell populations. Depending on the DRG neuron types that exist within the DRGs, the cellular response to pain is known to be different. Therefore, isolating a specific type of DRG neuron from a heterogeneous sample allows researchers to get a better understanding on its biochemistry and electrical signal on neurotransmission, which could be relevant for drug development.
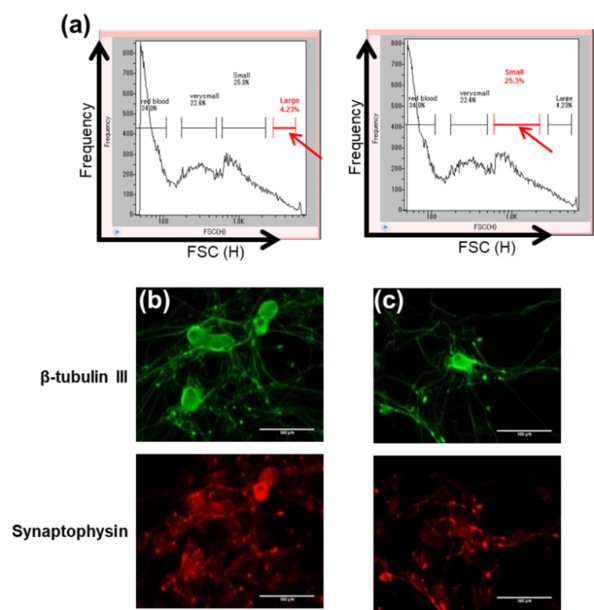
Fig. 5. (a) Scatter plots showing the two gates used for isolation of two separate cell populations (indicated by red arrows, named ‘large’ and ‘small’) from a heterogeneous DRGs sample by size on On-chip Sort. (b) Confirmation of nerve cells in the isolated ‘large’ (b) and ‘small’ (c) cell populations in (a) by staining of anti-β-tubulin III (green) and anti-Synaptophysin (red). The results are obtained in conjunction with Dr. Suzuki from Tohoku Institute of Technology, Japan.
DRGs collected from a 10-week old rat were dissociated by enzyme treatment and suspended in Neural Basal Medium prior to sorting. The pre-treated sample was sorted according to size (Fig. 5). Four groups of cells were observed from the size distribution, and two of these cell populations were separately isolated on On-chip Sort and cultured for five days. Both populations were fixed by paraformaldehyde, and stained with anti-β-tubulin III, which recognizes microtubule protein found in neurons, and anti-Synaptophysin, which recognizes neurons responsible for neurotransmission. Due to the anti-β-tubulin III and anti-Synaptophysin staining, both populations were confirmed to be viable nerve cells in DRGs. Therefore, these results reveal that On-chip Sort is capable of isolating DRG neurons by size while maintaining viability.
Induced pluripotent stem cells
In the field of regenerative medicine, differentiated somatic cells from embryonic stem cells or iPSCs are transplanted to patients. However, any remaining undifferentiated cells can become tumorous once they are implanted, obstructing the advancement in patient treatment. The removal of undifferentiated cells has been a challenge, and thus there is a need to efficiently remove the undifferentiated cells prior to transplantation. On-chip Sort effectively and efficiently removes rare undifferentiated cells present in the sample by a method called ‘multi-step negative sorting’.
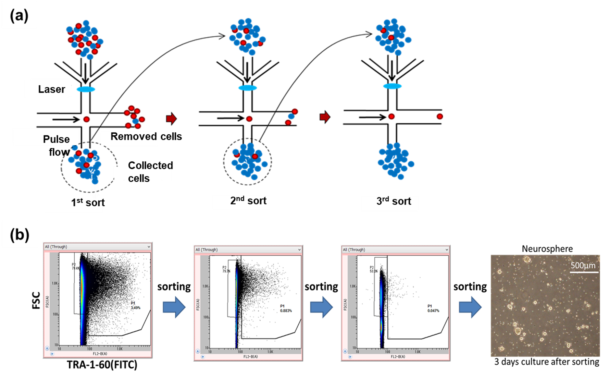
Fig. 6. (a) Principle of multi-step negative sorting. Red dots are undifferentiated cells and blue dots are differentiated cells. (b) Removal of undifferentiated cells from a sample of differentiated neural stem cells by multi-step negative sorting. The results are obtained in conjunction with Dr. Chuma from Institute for Frontier Medical Sciences, Kyoto University, Japan.
Multi-step negative sorting with On-chip Sort works by loading the sample (differentiated cells mixed with a rare population of undifferentiated cells) to the sample reservoir of the chip. Undifferentiated cells (non-target cells) are removed by formation of the pulse flows into the collection reservoir, while differentiated cells (target cells) flow into the waste reservoir located downstream of the chip. Differentiated cells are recovered and reloaded to the sample reservoir for further purification. This process is repeated until all the undifferentiated cells are removed (Fig. 6a). Fig. 6b shows the removal of ~4% of undifferentiated cells remaining in the sample containing cells that were differentiated to neural stem cells. Undifferentiated cells were fluorescently labeled with FITC-conjugated anti-TRA-1-60, and the fluorescence signals were used for the detection. We confirmed that all undifferentiated cells were removed after three sorts. We also cultured the sorted differentiated cells and found them viable (Fig. 6b).
Sperm cells
Sperm cells are known to be extremely sensitive to SICS, and a large number of sperm that are sorted using a conventional sorter are not suitable for use in in vitro fertilization (IVF). The effect of sperm sorting on On-chip Sort was evaluated using parameters such as motility, fertility and developmental ability. The overall procedure of the experiments is presented on Fig. 7. Oocytes and sperm were retrieved from C57BL/6L mice and sperm were pre-incubated in BSA-free Toyoda Yokoyama Hosi medium. Pre-incubated sperm was sorted using On-chip Sort with calcium-enhanced human tubal fluid (mHTF) as sheath fluid. IVF was performed on a dish with the collected sperm and retrieved oocytes in mHTF. In order to determine the developmental ability of the cultured two-cell embryos produced by IVF using collected sperm, embryo were transferred to Jcl:ICR mice.
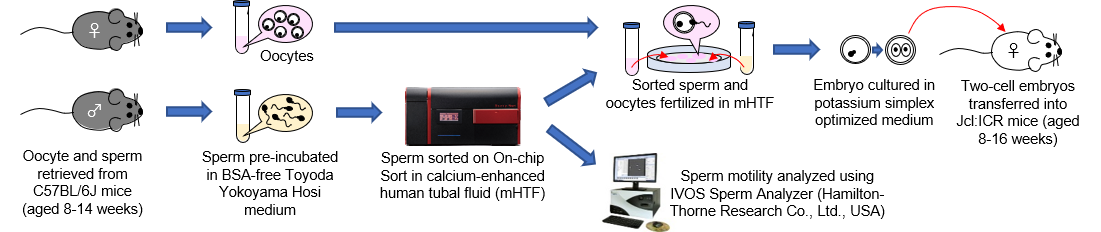
Fig. 7. Overall experimental procedure.
Sperm sorting on On-chip Sort was performed based on signals from forward scattered light (FSC) and side scattered light (SSC). The sorted sperm had lower motility than unsorted sperm (Fig. 8a), but the fertilization rate showed no significant difference between unsorted and sorted sperm (Fig. 8b). The sorted sperm were fertilized with oocytes, and the fertilized eggs developed normally up to the blastocyst stage in in vitro culture, which was comparable to those that were unsorted (Fig. 8c). Normal pups were obtained after embryo transfer, and birth rate was similar for both that used unsorted and sorted sperm (Fig. 8d). On-chip Sort has shown the mouse sperm isolation capability while maintaining normal fertilization and developmental ability.
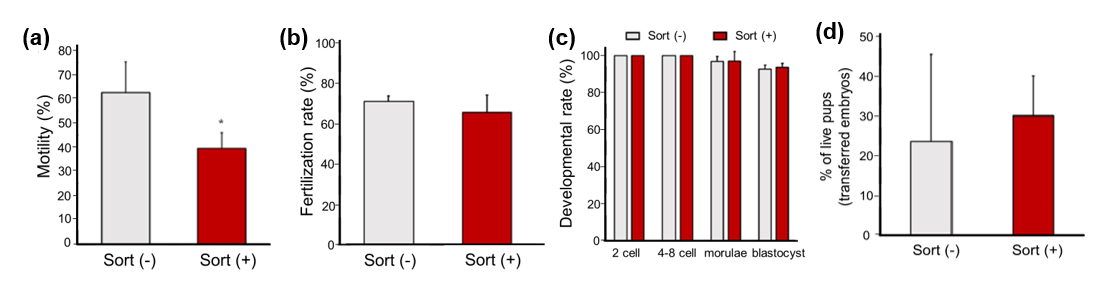
Fig. 8. (a) Motility of sperm with and without sorting. (b) Fertilization rate of sperm with and without sorting. (c) Developmental rate of sperm with and without sorting at several developmental stages. (d) Birth rate of live pup derived from unsorted and sorted sperm.
The data were obtained from Nakao, S., Takeo, T., Watanabe, H. et al. Successful selection of mouse sperm with high viability and fertility using microfluidics chip cell sorter. Sci. Rep. 10, 8862 (2020), under the license of Attribution 4.0 International (CC BY 4.0) https://creativecommons.org/licenses/by/4.0/.
Cell-cell complexes
On-chip Sort is able to isolate cells that are held together by weak bonds, even those that can easily be dissociated by pipetting. Fig. 9 shows the isolation of cell doublets (green and red cells) held together by weak intercellular interactions from a heterogeneous cell mixture.
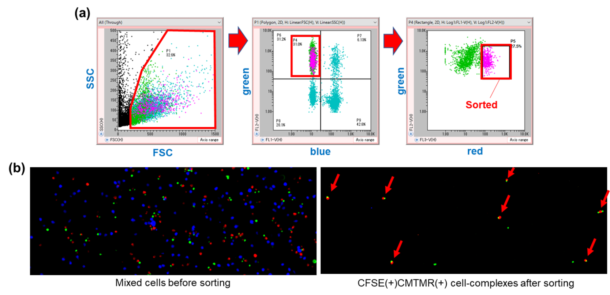
Fig. 9. (a) Sorting of green and red cell doublets held together by weak intercellular interactions from a heterogeneous sample. (b) Sample before (left) and after (right) sorting by On-chip Sort.